Save costs on your Sanger Sequencing kits - Affordable reactions
Are you in need of Sanger sequencing kits?
Are you planning to start your research with it?
Are you planning to start your research with it?
Sanger Sequencing is a fast, cost-effective method of sequencing small targeted regions of the genome. It is widely used to test for known familial variants, for validation of results obtained through NGS, and for some single gene sequencing. We offer affordable and high quality kits for the whole workflow of your Sanger Sequencing experiments.
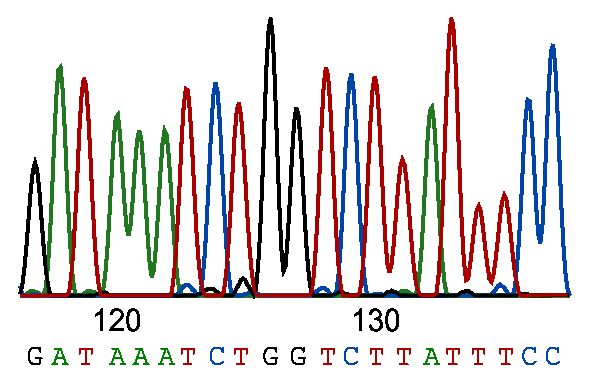
La secuenciación por el método de Sanger, utiliza la electroforesis capilar. Los pasos para el proceso de secuenciación de Sanger incluyen la fragmentación previa del ADN a secuenciar, seguido de la amplificación de estos fragmentos mediante la reacción en cadena de la polimerasa (PCR). A continuación, cada fragmento acabado en didesoxinucleótido está marcado fluorescentemente, de manera que cada didesoxinucleótido (ddATP, ddCTP, ddGTP, ddTTP) implica un color de fluoróforo distinto.
Los pasos importantes en la secuenciación de Sanger son los pasos de purificación y el uso de productos de alta calidad. Esto eliminará cualquier interferencia y aumentará la intensidad y el espaciado de los picos.
Desde Isogen Life Science ofrecemos kits para los distintos pasos del flujo de trabajo.
We offer kits from each of the workflow steps:
The purification step for PCR products, such as DNA polymerase, dNTPs or primers can be performed with use of magnetic beads. These beads are less time consuming and more inexpensive than the common used spin columns. They are adaptable to high throughput research, recover high amounts of DNA, form a uniform suspension and are suitable for a DNA sequencing workflow.
ADS PCR Cleaning Magnetic Beads: Our PCR cleaning beads enable you the following:
For more information about it, have a look at the brochure here!
Also, we offer other kits and reagents like:

|
In the Sanger DNA sequencing reaction, linear amplifications are labelled with one nucleotide difference with a ddNTP to terminate the chain. For this step we offer the SupreDye Cycle Sequencing Kit, which provides high performance, increase in peaks and longer read lengths. We also offer cycle sequence kits for high G-templates: SupreDye dGTP Sequencing Kit.
Achieve high performance and cost-effective quality results with our affordable cycle sequencing kits: |
Thereafter there is need for a sequence clean up to remove contaminants and unused ddNTPs. Different ways for this purification can be used. The ADS BD-XT Purification Kit is fast, reliable, has a simple work flow and does not need a follow up washing step. The ADS Sequence Reaction Cleaning Beads is another option for the purification, as the contaminants will bind to the beads, it can easily be removed. This method is ideal for DNA sequencing, requires a minimal amount of washing steps and is cost-effective.
Make sure contaminants and unincorporated ddNTP dyes from your sample are removed before sample loading. Discover our solutions:
After the polymerase chain reaction the sample can be loaded on capillary electrophoresis. Depending on the purification method, the sample can be directly loaded or needs resuspension. When the sample is purified with ethanol the sample is often resuspended with formamide. For the optimal signal intensity and stability TruPure formamide can be used. Another method is the use of polymers. The PwrPOP polymers are effective for separation of extension products. This is a cost-effective solution and creates even peak separation and long reads.
After extensive usage of the capillary electrophoresis, there is a need for regeneration, which restores the performance of the capillary array. If this is not performed after approximately 500 runs the lifetime of the capillary array is reduced. The replacement of the capillary array is more expensive than the regular maintenance. The need for maintenance is recognisable on the uneven peak space and short reading lengths. For the regeneration we offer the ADS Capillary Regeneration Kit. It is easy to implement, has an on instrument protocol and a short hands-on time and there is no need for capillary disassembly.
Discover more about our Capillary array regeneration kit:

Have a look at our solutions for genomics, cell biology, diagnostics and proteins. We offer intruments, reagents, kits and consumables for your research!
+31 30 6880771
support@isogen-lifescience.com
Veldzigt 2A
3454 PW De Meern
