THOR technology can be used to generate individual and pooled 3’ mRNA-seq, or full-length individual libraries from ultra-low input RNA or single cells. The technology is template-switch-, and ligation-free, and employs a unique THOR reaction. The THOR reaction is initiated at oligo(dT ) primed poly(A) tails introducing a T7 promoter sequence to all 3’ ends of transcripts. The resulting structure allows swift amplification of antisense RNA directly from mRNA templates.
To discover more about the technology, you can have a look at the poster: "Unparalleled sensitivity in single cell sequencing. The THOR technology enables then to achieve the best sensitivity when performing scRNA Seq in low input samples.
- True single-cell expression profiling
- High resolution for ultra-low input
- Low input scRNA-Seq samples – when you want to analyze from 1-100 cells
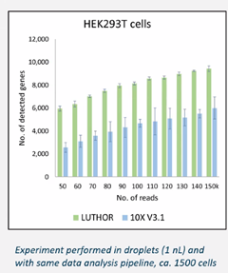
- Very high gene detection rate
- Categorize cells by RNA expression. Compared to 10x, classification between subpopulations is now possible with LUTHOR