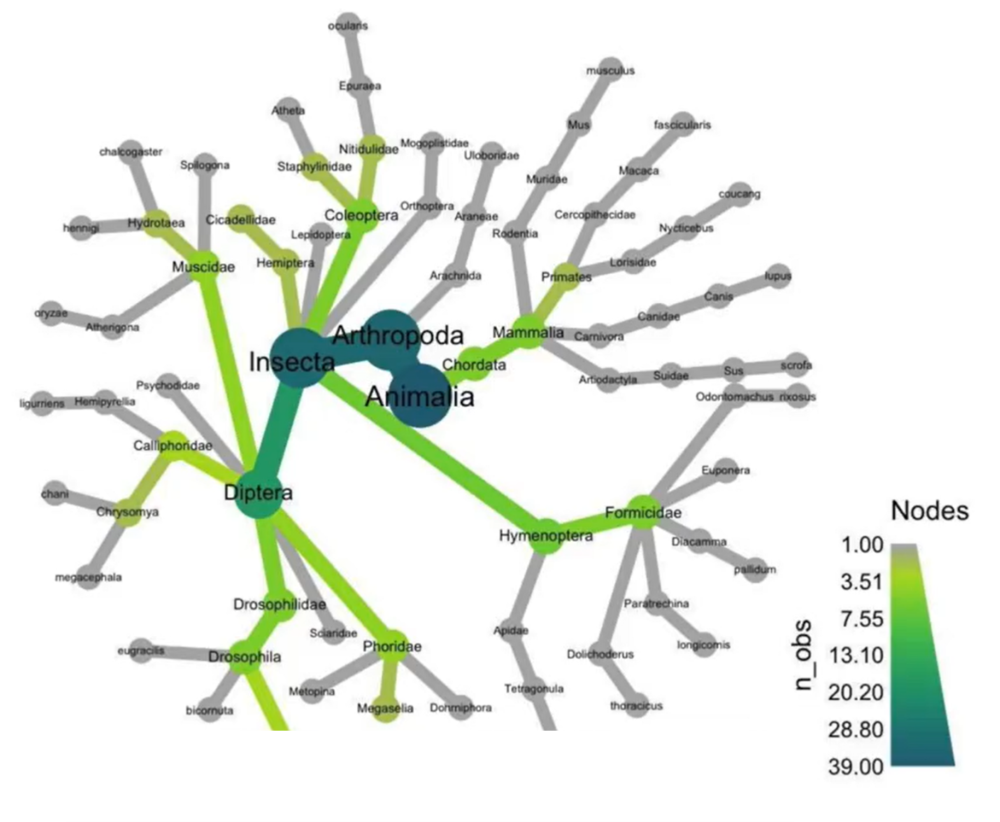
Key findings: 313bp barcode amplified in just 22 minutes!
Flies were captured with the robots in the field and were analyzed later. The analysis was performed by sequencing fly poop, and therefore gut content samples.
The protocol was based on the workflow that the Museum für Naturkunde in Berlin performed. The PCR amplification step for the COI fragment took only 22 minutes using the NextGenPCR thermocycle. Therefore, it was possible to barcode 250 fly specimens in just 6 hours, which would have taken significantly longer using the traditional method. Moreover, it uses 80x less energy.