PureIT ExoZAP PCR Cleanup
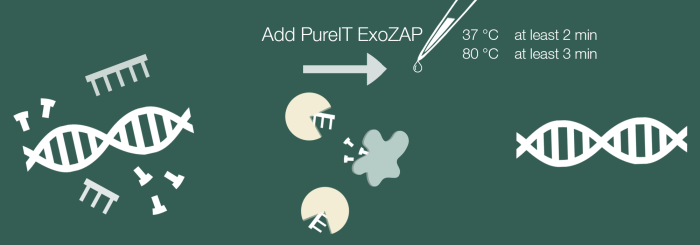
Amplified PCR products can cause troubles in downstream applications due to excess of primers, ssDNA and/or dNTPs still being present. Treating your amplified PCR products with PureIT ExoZAP PCR Cleanup, an enzymatic PCR clean up, will make your samples ready for applications such as sequencing.
Features PCR Cleanups
- One-step PCR Cleanup
- Fast protocol, only 5 minutes
- No need for spin columns or magnetic beads
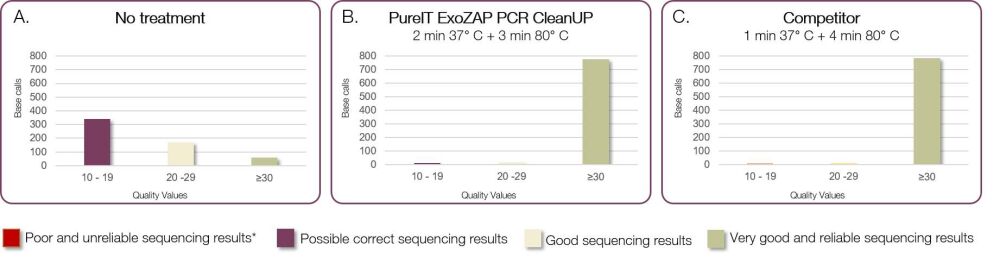
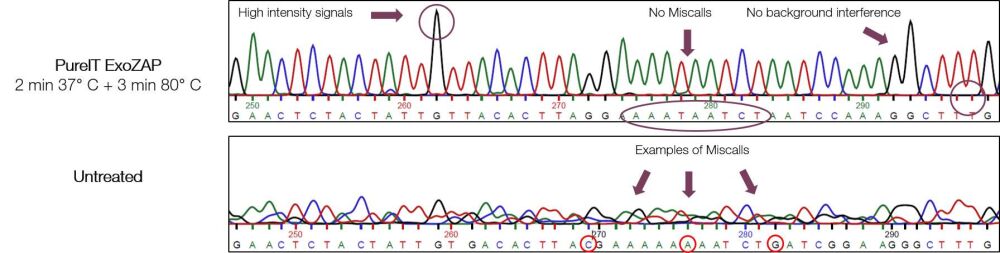
- Treatment improves downstream applications such as DNA sequencing and SNP analysis
- Minimal loss of PCR products during treatment
Do you want to avoid downstream challenges when running your PCR?
Contact us for more information about ExoZap
We will get back to you in 1 to 2 working days